Back to article: Towards a better understanding of the neuro-developmental role of autophagy in sickness and in health
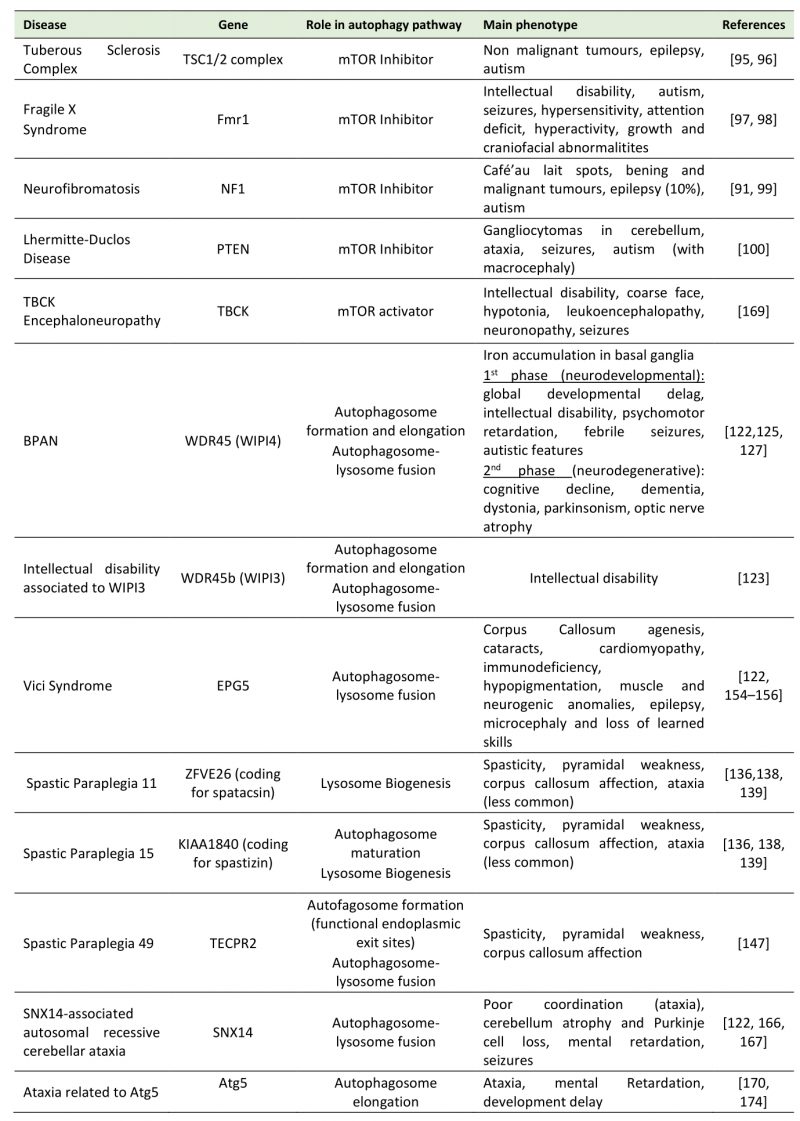
TABLE 1. Main neurodevelopmental disorders where mutations in autophagy proteins impacts disease.
91. Liberti M V., and Locasale JW (2016). The Warburg Effect: How Does it Benefit Cancer Cells? Trends Biochem Sci 41(3): 211–218. 10.1016/j.tibs.2015.12.001
95. Fleming A, and Rubinsztein DC (2020). Autophagy in Neuronal Development and Plasticity. Trends Neurosci 43(10): 767–779. 10.1016/j.tins.2020.07.003
96. Kuijpers M, and Haucke V (2021). Neuronal autophagy controls the axonal endoplasmic reticulum to regulate neurotransmission in healthy neurons. Autophagy 17(4): 1049-1051. 10.1080/15548627.2021.1893569
97. Waites CL, Craig AM, and Garner CC (2005). Mechanisms of vertebrate synaptogenesis. Annu Rev Neurosci 28(1): 251–74. 10.1146/annurev.neuro.27.070203.144336
98. Tang G, Gudsnuk K, Kuo S-H, Cotrina ML, Rosoklija G, Sosunov A, Sonders MS, Kanter E, Castagna C, Yamamoto A, Yue Z, Arancio O, Peterson BS, Champagne F, Dwork AJ, Goldman J, and Sulzer D (2014). Loss of mTOR-dependent macroautophagy causes autistic-like synaptic pruning deficits. Neuron 83(5): 1131–43. 10.1016/j.neuron.2014.07.040
99. Association AP (2013). Diagnostic and statistical manual of mental disorders., 5th ed. American Psychiatric Association.
100. Bourgeron T (2009). A synaptic trek to autism. Curr Opin Neurobiol 19(2): 231–234. 10.1016/j.conb.2009.06.003
122. Wang T, Guo H, Xiong B, Stessman HAF, Wu H, Coe BP, Turner TN, Liu Y, Zhao W, Hoekzema K, Vives L, Xia L, Tang M, Ou J, Chen B, Shen Y, Xun G, Long M, Lin J, Kronenberg ZN, Peng Y, Bai T, Li H, Ke X, Hu Z, Zhao J, Zou X, Xia K, and Eichler EE (2016). De novo genic mutations among a Chinese autism spectrum disorder cohort. Nat Commun 7(13): 1–10. 10.1038/ncomms13316
123. Bateup HS, Johnson CA, Denefrio CL, Saulnier JL, Kornacker K, and Sabatini BL (2013). Excitatory / Inhibitory Synaptic Imbalance Leads to Hippocampal Hyperexcitability in Mouse Models of Tuberous Sclerosis. Neuron 78(3): 510–522. 10.1016/j.neuron.2013.03.017
125. Talos DM, Sun H, Kosaras B, Joseph A, Folkerth RD, Poduri A, Madsen JR, Black PM, and Jensen FE (2012). Altered inhibition in tuberous sclerosis and type IIb cortical dysplasia. Ann Neurol 71(4): 539–551. 10.1002/ana.22696
127. D'Hulst C, Heulens I, Brouwer JR, Willemsen R, De Geest N, Reeve SP, De Deyn PP, Hassan BA, and Kooy RF (2009). Expression of the GABAergic system in animal models for fragile X syndrome and fragile X associated tremor/ataxia syndrome (FXTAS). Brain Res 1253: 176–183. 10.1016/j.brainres.2008.11.075
131. Yoganathan S, Arunachal G, Sudhakar SV, Rajaraman V, Thomas M, and Danda S (2016). Beta Propellar Protein-Associated Neurodegeneration: A Rare Cause of Infantile Autistic Regression and Intracranial Calcification. Neuropediatrics 47(2): 123–127. 10.1055/s-0035-1571189
136. Hor CHH, and Tang BL (2019). Beta-propeller protein-associated neurodegeneration (BPAN) as a genetically simple model of multifaceted neuropathology resulting from defects in autophagy. Rev Neurosci 30(3): 261–277. 10.1515/revneuro-2018-0045
138. Behrends C, Sowa ME, Gygi SP, and Harper JW (2010). Network organization of the human autophagy system. Nature 466(7302): 68–76. 10.1038/nature09204
147. Vantaggiato C, Crimella C, Airoldi G, Polishchuk R, Bonato S, Brighina E, Scarlato M, Musumeci O, Toscano A, Martinuzzi A, Santorelli FM, Ballabio A, Bresolin N, Clementi E, and Bassi MT (2013). Defective autophagy in spastizin mutated patients with hereditary spastic paraparesis type 15. Brain 136(10): 3119–3139. 10.1093/brain/awt227
154. Branchu J, Boutry M, Sourd L, Depp M, Leone C, Corriger A, Vallucci M, Esteves T, Matusiak R, Dumont M, Muriel MP, Santorelli FM, Brice A, El Hachimi KH, Stevanin G, and Darios F (2017). Loss of spatacsin function alters lysosomal lipid clearance leading to upper and lower motor neuron degeneration. Neurobiol Dis 102: 21–37. 10.1016/j.nbd.2017.02.007
155. Denora PS, Smets K, Zolfanelli F, Groote CC De, Casali C, Deconinck T, Sieben A, Gonzales M, Zuchner S, Darios F, Peeters D, Brice A, Malandrini A, De Jonghe P, Santorelli FM, Stevanin G, Martin JJ, and El Hachimi KH (2016). Motor neuron degeneration in spastic paraplegia 11 mimics amyotrophic lateral sclerosis lesions. Brain 139(6): 1723–1734. 10.1093/brain/aww061
156. Oz-Levi D, Ben-Zeev B, Ruzzo EK, Hitomi Y, Gelman A, Pelak K, Anikster Y, Reznik-Wolf H, Bar-Joseph I, Olender T, Alkelai A, Weiss M, Ben-Asher E, Ge D, Shianna K V, Elazar Z, Goldstein DB, Pras E, and Lancet D (2012). Mutation in TECPR2 reveals a role for autophagy in hereditary spastic paraparesis. Am J Hum Genet 91(6): 1065–1072. 10.1016/j.ajhg.2012.09.015
166. Byrne S et al. (2016). EPG5-related Vici syndrome: A paradigm of neurodevelopmental disorders with defective autophagy. Brain 139(3): 765–781. 10.1093/brain/awv393
167. Cullup T et al. (2013). Recessive mutations in EPG5 cause Vici syndrome, a multisystem disorder with defective autophagy. Nat Genet 45(1): 83–87. 10.1038/ng.2497
169. Tian Y, Li Z, Hu W, Ren H, Tian E, Zhao Y, Lu Q, Huang X, Yang P, Li X, Wang X, Kovács AL, Yu L, and Zhang H (2010). C. elegans Screen Identifies Autophagy Genes Specific to Multicellular Organisms. Cell 141(6): 1042–1055. 10.1016/j.cell.2010.04.034
170. Wang Z, Miao G, Xue X, Guo X, Yuan C, Wang Z, Zhang G, Chen Y, Feng D, Hu J, and Zhang H (2016). The Vici Syndrome Protein EPG5 Is a Rab7 Effector that Determines the Fusion Specificity of Autophagosomes with Late Endosomes/Lysosomes. Mol Cell 63(5): 781–795. 10.1016/j.molcel.2016.08.021
174. Akizu N et al. (2015). Biallelic mutations in SNX14 cause a syndromic form of cerebellar atrophy and lysosome-autophagosome dysfunction. Nat Genet 47(5): 528–534. 10.1038/ng.3256